
Research
Neurons in the brain have a remarkable capacity to undergo changes in function and structure throughout life. How does sensory experience alter neuronal activity and ultimately triggers behavioral adaptations? We address this question by studying the role of neuronal biochemical signaling dynamics in experience dependent plasticity. We use advanced microscopy, gene editing and in vivo imaging to unravel protein signaling dynamics and neuronal activity in awake behaving mice.
Experience dependent dynamics of activity dependent transcription in intact cortical circuits
One major pathway which induces long-lasting modulation of neuronal properties are activity dependent transcription factors, vital for experience dependent plasticity and long-term memory. They have been hypothesized to convert short-lasting Calcium signaling via neuronal and synaptic activity into long-lasting transcriptional and translation, eventually leading to changes in functional properties of neuronal circuits. We study dynamics of neuronal signaling during ongoing sensory experience by using biosensors adapted for in vivo two-photon fluorescence lifetime imaging (2pFLIM). This allows high resolution quantitative imaging of distinct signaling events during sensory stimulation and experience. We have established a new method that allows to measure the activity of CREB, a well-known evolutionary conserved transcription factor which is important for LTP and memory. We have made new biosensors and developed an approach to simultaneously monitor neuronal activity and CREB in single cell resolution in awake behaving mice. This unique platform will allow us to dissect the coupling between neuronal activity and transcription and its underlying mechanisms, uncovering the biochemical signaling motifs which mediate experience deponent plasticity of neuronal circuits.


In vivo 2pFLIM of a mouse which is adapting to run on a rotating disc. The images show ongoing changes in calcium activity (upper panel) and CREB activity (lower panel) during a 60 minute imaging session.
Synapse to nucleus communication in intact cortical circuits
In order to faithfully respond and adapt to changes in the environment, synaptic activity must be integrated and communicated to the nucleus. Previous work has identified that strong synaptic activity in vitro can induce translocation of numerous signaling proteins from synapses to the nucleus. However, the extent and mechanisms of in vivo translocation and signaling to the nucleus following physiological patterns of synaptic activity are scarce. In this project, we will examine if and how specific synaptic/nuclear proteins undergo nuclear translocation and their role in experience-dependent sensory stimuli. To explore this, we will utilize newly developed gene editing tools, which allow genomic labeling using CRISPR/Cas9 in single neurons in the brain. This technique will enables to directly monitor innate intracellular signaling dynamics which underlie communication between synapse to nucleus during experience dependent plasticity.
Synapse
Nucleus



In utero electroporation of dividing cortical cells allows CRISPR Cas9 endogenous protein visualization in the brain. Images of HA-tagged CaMKIIg at synapses and CREB at the nucleus in cortical neurons.
Dynamics of epigenetics regulation in the brain
The neuronal genome undergoes widespread epigenetic regulation which is critical for normal brain development and adult brain plasticity. The importance of this modulation is demonstrated by many devastating neurological disorders and cognitive deficiencies which have been genetically associated with epigenetic factors and transcription modulators. We will utilize and develop new experimental approaches to allow dynamic visualization of epigenetic signaling in the brain. This will allow us to directly link ongoing epigenetic, neuronal activity and behavioral states, and to understand their physiological role. Ultimately, deciphering epigenetic signaling rules at the healthy brain will enable shed light on how its dysfunction leads to devastating cognitive decline and neurological disorders.
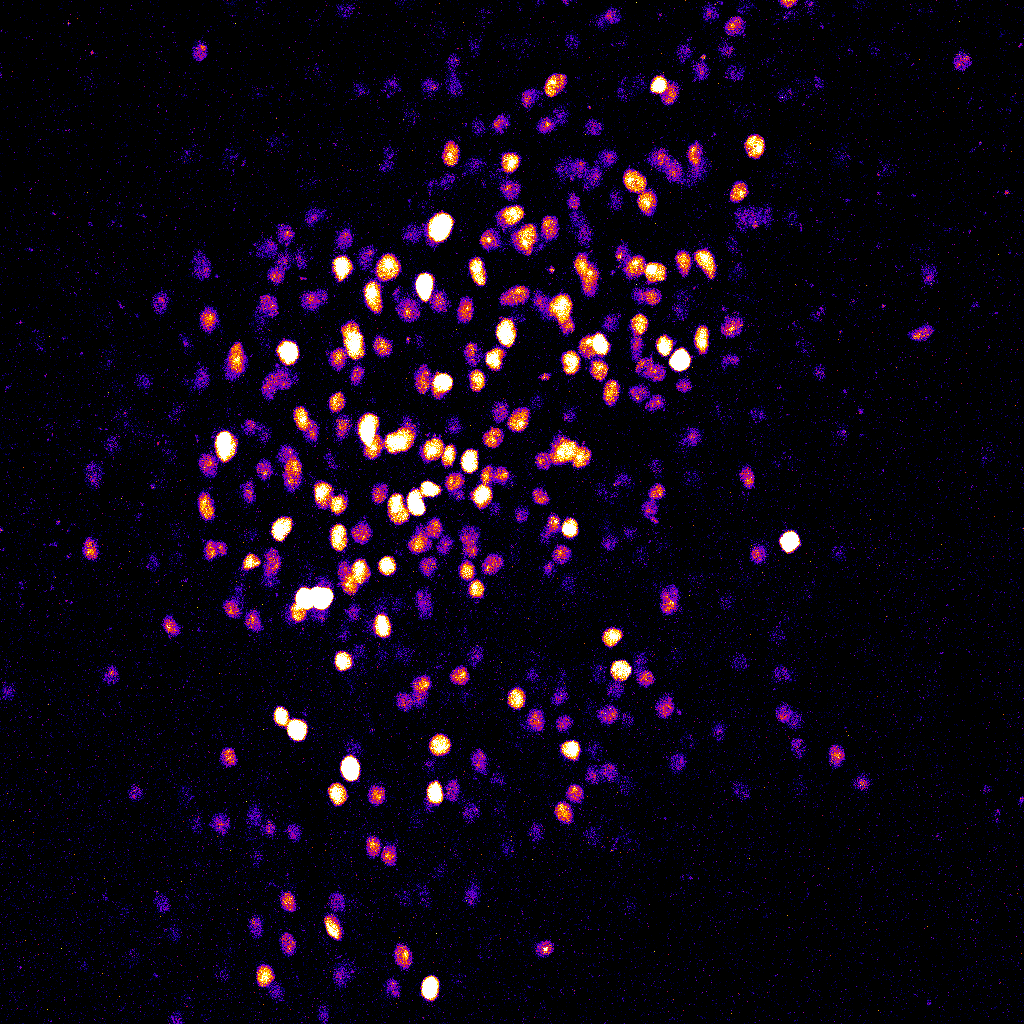
MeCP2-GFP expression in L2/3 neurons

In vivo imaging of synaptic activity in a L2/3 cell expressing GCaMP/CyRFP in the motor cortex
Development of new techniques for monitoring and manipulation of neuronal signaling in living mice
We are working to develop new approaches to dissect neuronal signaling:
-New 2pFLIM biosensors for important signaling proteins.
-Methods for tagging of endogenous proteins in the brain using CRISPR/Cas9..
-Manipulation of protein function and expression via optogenetic control of important signaling pathways.
- Labeling and manipulation of signaling protein in non-genetic animal models.
